Matrix Multiplication in Polars: A 2026 Guide to Distance Computation
Every ML engineer has been here: you have two sets of embeddings, and you need to find which items in set B are most similar to each item in set A. Maybe you’re building a recommendation system, a semantic search feature, or a simple kNN classifier.
In NumPy, this is a one-liner:
similarities = query_embeddings @ corpus_embeddings.TThe @ operator performs matrix multiplication. For normalized vectors, this computes cosine similarity. It’s fast because NumPy delegates to highly optimized BLAS libraries (OpenBLAS, MKL, or Apple’s Accelerate).
But what if your data lives in a DataFrame? What if you need to filter by category, date range, or other business logic before computing similarities? This is where things get interesting.
The Question
Can Polars perform matrix multiplication efficiently enough to replace NumPy for distance matrix computation?
The short answer: No—but it doesn’t have to.
The longer answer: Polars excels at data wrangling, filtering, and transformations. NumPy excels at dense linear algebra. The winning strategy is to combine them.
What About Approximate Nearest Neighbor (ANN) Engines?
Libraries like FAISS, Annoy, and ScaNN are designed for billion-scale search where exact computation is infeasible. They build index structures that enable sub-linear query time at the cost of some recall.
This article focuses on exact computation for datasets that fit in memory—tens of thousands to low millions of vectors. If your corpus has 100 million+ vectors, you need an ANN engine. If it has 50K vectors and you need exact results with complex filtering, read on.
The Approaches
I benchmarked five approaches:
| Approach | Description |
|---|---|
| NumPy | Pure matrix multiplication with @ operator |
| Pandas | Cross-merge + apply for dot product |
| Polars DataFrame | Cross-join + map_elements for dot product |
| Polars LazyFrame | Same as above with lazy evaluation |
| Polars + NumPy | Use Polars for filtering, NumPy for math |
What About polars-ds and Other Plugins?
Before benchmarking, I investigated the Polars plugin ecosystem. polars-ds (Polars for Data Science) caught my attention—it provides native Rust implementations of:
query_radius_ptwise: Find all neighbors within radius rarr_sql2_dist/list_sql2_dist: Squared L2 distance between arrays/lists- KNN queries with k-d tree indexing (in development)
However, the polars-ds documentation explicitly states:
“Due to limitations, this currently doesn’t preserve the index, and is not fast when k or dimension of data is large.”
The library targets datasets up to 2-3 million rows and isn’t designed to compete with BLAS-optimized matrix multiplication for bulk distance computation. It’s excellent for per-row operations and small-scale KNN, but not for the distance matrix use case we’re exploring.
Similarly, polars-ols provides fast linear regression using Rust + LAPACK, but doesn’t expose general matrix multiplication.
Bottom line: The plugin ecosystem is growing, but for serious linear algebra, you still need NumPy (or JAX/PyTorch).
Implementation Details
NumPy: The Baseline
def numpy_topk(query: np.ndarray, corpus: np.ndarray, k: int) -> np.ndarray:
# Matrix multiplication: O(n_query * n_corpus * n_dims)
similarities = query @ corpus.T
# Efficient top-k selection: O(n_query * n_corpus)
partitioned = np.argpartition(-similarities, k, axis=1)[:, :k]
# Sort within top-k: O(n_query * k * log(k))
rows = np.arange(len(query))[:, None]
top_k_similarities = similarities[rows, partitioned]
sorted_within_k = np.argsort(-top_k_similarities, axis=1)
return partitioned[rows, sorted_within_k]Key insight: np.argpartition is O(n) for selecting top-k elements, much faster than full sorting.
Polars Pure: The DataFrame Way
Since Polars 1.8+, list columns support native arithmetic, enabling a cleaner approach:
def polars_df_topk(query: np.ndarray, corpus: np.ndarray, k: int) -> np.ndarray:
query_df = pl.DataFrame({
"query_id": range(len(query)),
"query_vec": query.tolist()
})
corpus_df = pl.DataFrame({
"corpus_id": range(len(corpus)),
"corpus_vec": corpus.tolist()
})
# Cross join creates all query × corpus pairs
joined = query_df.join(corpus_df, how="cross")
# Native list arithmetic: element-wise multiply, then sum
result = joined.with_columns(
(pl.col("query_vec") * pl.col("corpus_vec")).list.sum().alias("similarity")
)
# Get top-k per query
return result.sort(["query_id", "similarity"], descending=[False, True]) \
.group_by("query_id", maintain_order=True).head(k)This is cleaner than using map_elements (which drops into Python), but memory is still the killer. The cross-join materializes all n_query × n_corpus pairs, each carrying the full vectors.
Polars + NumPy Hybrid: Best of Both Worlds
def polars_numpy_topk(
query: np.ndarray,
corpus: np.ndarray,
k: int,
metadata: dict
) -> np.ndarray:
# Use Polars for complex filtering
corpus_df = pl.DataFrame({
"corpus_id": range(len(corpus)),
"category": metadata["corpus_category"],
"date": metadata["corpus_date"]
})
filtered = corpus_df.filter(
pl.col("category").is_in(metadata["included_categories"])
& (pl.col("date") >= metadata["date_start"])
& (pl.col("date") <= metadata["date_end"])
)
# Get filtered indices and subset the matrix
corpus_indices = filtered["corpus_id"].to_numpy()
corpus_filtered = corpus[corpus_indices]
# Use NumPy for the math
similarities = query @ corpus_filtered.T
top_k_local = np.argpartition(-similarities, k, axis=1)[:, :k]
# Map back to original indices
return corpus_indices[top_k_local]Benchmark Results
Test Configuration
- Dimensions: 100, 1,000, 3,000 (matching real embedding sizes like Gemini’s 3,072-dim text embeddings)
- Matrix sizes: 500 queries × 20K corpus, 1K × 50K, 2K × 100K
- Top-k: 1, 5, 10
- With/without filtering: Category membership + date range
Execution Time
The difference is stark. For a small dataset (100 queries × 2,000 corpus, 100 dimensions):
| Approach | Time | Slowdown vs NumPy |
|---|---|---|
| NumPy | 0.002s | 1× |
| Pandas | 0.45s | 250× |
| Polars DF (native) | 0.13s | 75× |
| Polars LazyFrame | 0.13s | 70× |
| Polars + NumPy | 0.002s | 1× |
When dimensions increase to 1,000 (matching typical embedding sizes):
| Approach | Time | Memory |
|---|---|---|
| NumPy | 0.002s | 400 MB |
| Pandas | 0.46s | 410 MB |
| Polars DF (native) | 2.0s | 7.8 GB |
| Polars LazyFrame | 1.1s | 6.4 GB |
| Polars + NumPy | 0.002s | 360 MB |
For pure matrix operations on 1000-dim vectors, NumPy is 900× faster than pure Polars—and that’s with native list operations. The slowdown comes from:
- Memory explosion: Cross-join stores all pairs with full vectors
- List overhead: Each cell is a Python list, not a contiguous array
- No BLAS: Even native ops don’t use SIMD or cache optimization
Memory Usage
The DataFrame approaches create a row for every query-corpus pair. With 100 × 2,000 = 200,000 pairs at 1,000 dimensions:
| Approach | Peak Memory | Memory Pattern |
|---|---|---|
| NumPy | 390 MB | Creates similarity matrix only |
| Pandas | 417 MB | Stores pairs as rows |
| Polars DF | 7.8 GB | All pairs + list column overhead |
| Polars Lazy | 6.4 GB | Slight optimization |
| Polars + NumPy | 401 MB | Same as NumPy |
At larger scales, pure Polars becomes infeasible. A 2,000 × 100,000 matrix (200M pairs) completes in 1.4s with NumPy but would take hours with pure Polars.
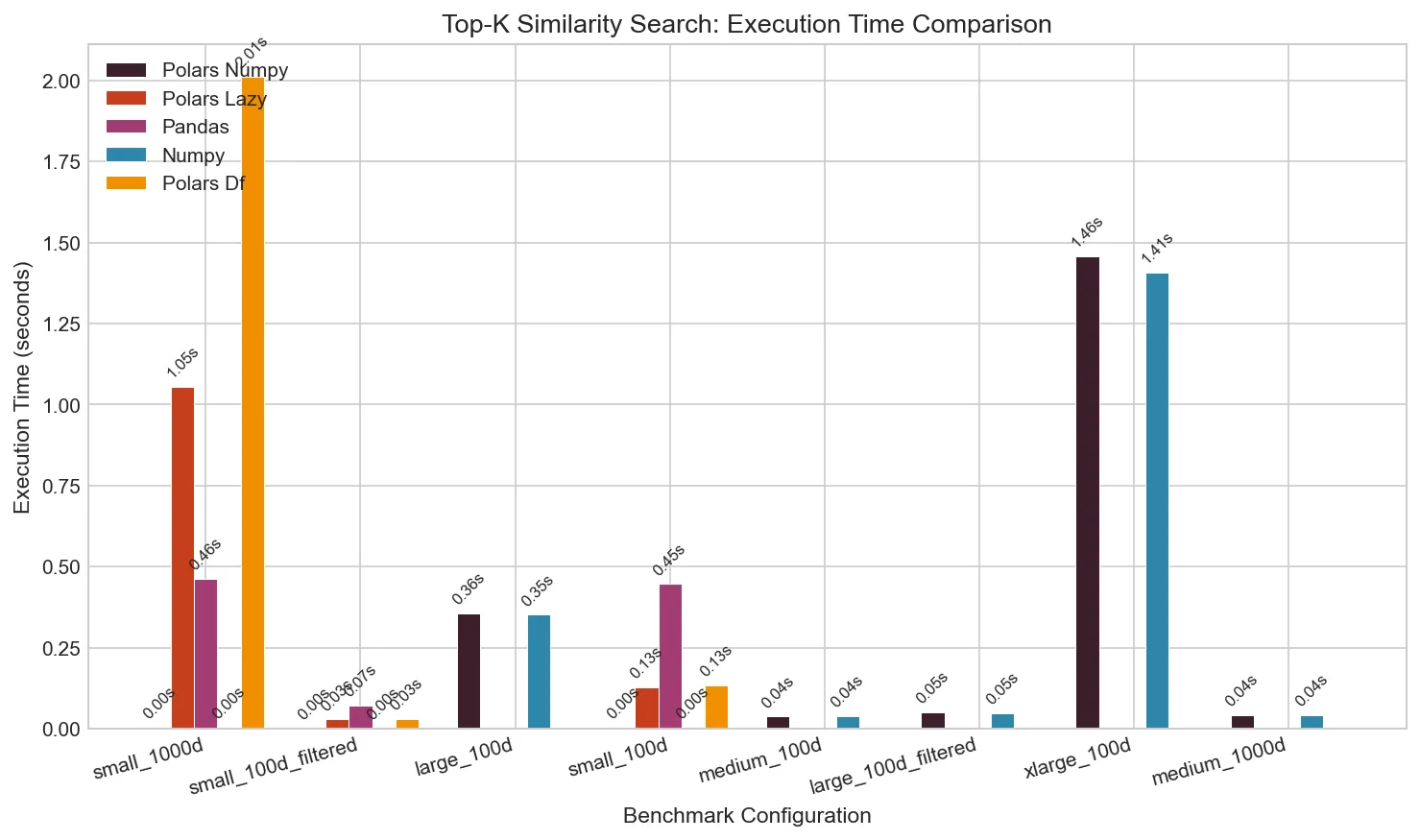
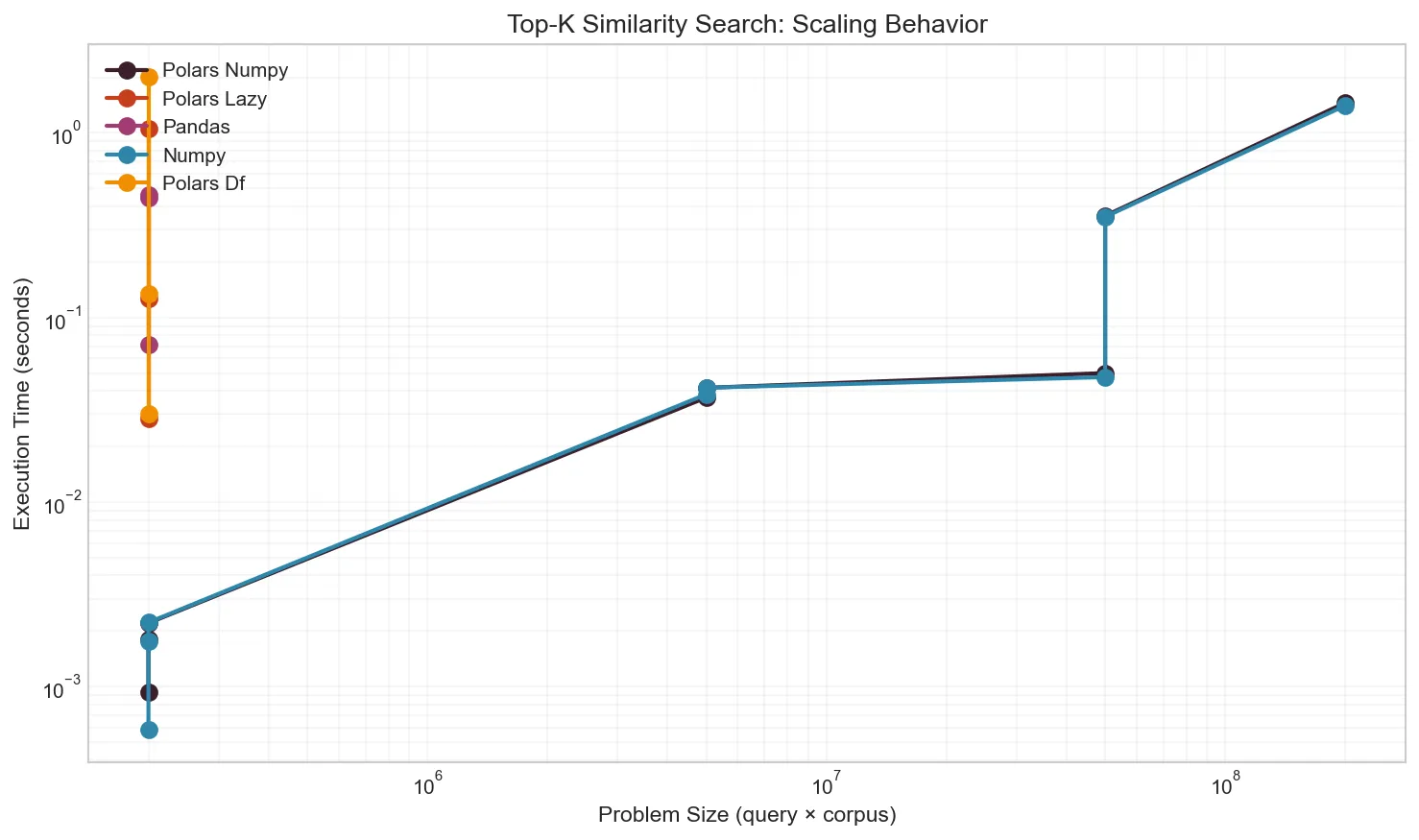
Effect of Filtering
When you need to filter the corpus before computing similarities, Polars shines:
# Filter 70% of corpus by category and date range
corpus_df.filter(
pl.col("category").is_in(allowed_categories)
& (pl.col("date").is_between(start, end))
)This filtering happens at native speed, often reducing the corpus by 50-90%. Then NumPy handles the smaller matrix multiplication.
When to Use What
flowchart TD
A[Distance Matrix Computation] --> B{Corpus size?}
B -->|> 1M vectors| C[Use ANN engine<br/>FAISS, ScaNN, etc.]
B -->|< 1M vectors| D{Need filtering?}
D -->|No| E[Pure NumPy]
D -->|Yes| F{Filter complexity?}
F -->|Simple| G[NumPy with boolean mask]
F -->|Complex| H[Polars filter → NumPy matmul]Use Pure NumPy When:
- No filtering needed, or filtering is simple enough for boolean masks
- You’re already working with numpy arrays
- Maximum performance is critical
Use Polars + NumPy Hybrid When:
- Complex filtering on multiple columns
- Date range queries
- Category membership
- Data originates from Parquet/CSV/database
- You want readable, maintainable code
Use Pure Polars When:
- Correctness matters more than speed
- Prototype/exploration phase
- Very small datasets (< 1000 pairs)
Avoid Pandas for This:
- The
.apply()pattern is slow - Memory overhead is significant
- Polars is strictly better for DataFrame operations
Reproducing These Results
The benchmark script uses uv for dependency management:
# Install uv if needed
curl -LsSf https://astral.sh/uv/install.sh | sh
# Run quick test
uv run benchmark.py --quick
# Run full benchmarks
uv run benchmark.py --output results/local
# Run in Docker with resource constraints
./run_benchmarks.shThe script automatically monitors CPU and memory usage, generates visualizations, and saves results to JSON.
Key Takeaways
-
NumPy’s matrix multiplication is unbeatable. The
@operator uses decades of BLAS optimization. Don’t try to replicate it in pure Python. -
Polars excels at what it’s designed for. Filtering, grouping, aggregating—these operations are fast. Use them.
-
Combine tools strategically. The hybrid Polars + NumPy approach gives you:
- Readable, expressive filtering with Polars
- Blazing-fast matrix math with NumPy
- Memory efficiency by filtering before computing
-
Avoid cross-join patterns for large matrices. Creating millions of row pairs kills memory and performance.
-
Consider your data’s lifecycle. If your data lives in Parquet files and needs complex transformations, Polars is the right entry point. If it’s already in numpy arrays, stay there.
Appendix: Full Benchmark Code
The complete benchmark script is available at benchmark.py. It includes:
- All five implementations
- Resource monitoring with psutil
- Publication-quality chart generation
- Docker wrapper for controlled environments
#!/usr/bin/env python3
# /// script
# requires-python = ">=3.12"
# dependencies = [
# "polars>=1.0.0",
# "numpy>=2.0.0",
# "pandas>=2.2.0",
# "matplotlib>=3.9.0",
# "psutil>=6.0.0",
# "rich>=13.0.0",
# ]
# ///
# Run with: uv run benchmark.pyFeel free to adapt it for your specific use case.